注意
前往結尾以下載完整範例程式碼。或透過 JupyterLite 或 Binder 在您的瀏覽器中執行此範例
物種分佈建模#
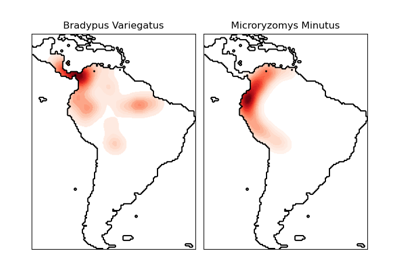
為物種的地理分佈建立模型是保護生物學中的一個重要問題。在此範例中,我們根據過去的觀測結果和 14 個環境變數,為兩種南美哺乳動物的地理分佈建立模型。由於我們只有正例(沒有不成功的觀測結果),我們將此問題視為密度估計問題,並使用 OneClassSVM 作為我們的建模工具。資料集由 Phillips 等人 (2006) 提供。如果可用,此範例會使用 basemap 來繪製南美的海岸線和國界。
這兩個物種是
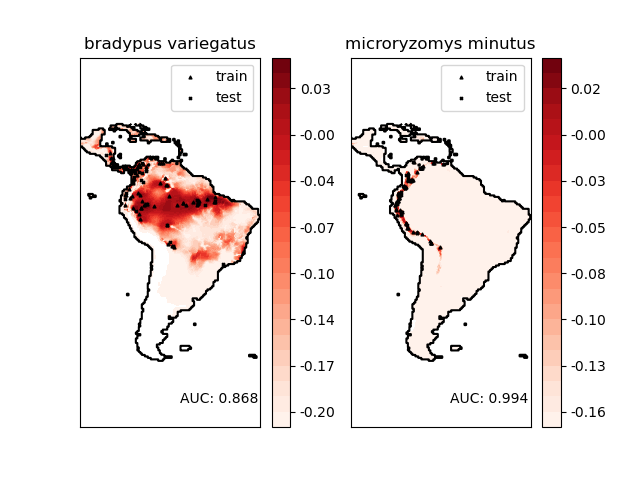
Bradypus variegatus,棕喉樹懶。
Microryzomys minutus,也稱為森林小稻鼠,一種生活在秘魯、哥倫比亞、厄瓜多、秘魯和委內瑞拉的囓齒動物。
參考文獻#
「物種地理分佈的最大熵建模」 S. J. Phillips、R. P. Anderson、R. E. Schapire - Ecological Modelling,190:231-259,2006。
________________________________________________________________________________
Modeling distribution of species 'bradypus variegatus'
- fit OneClassSVM ... done.
- plot coastlines from coverage
- predict species distribution
Area under the ROC curve : 0.868443
________________________________________________________________________________
Modeling distribution of species 'microryzomys minutus'
- fit OneClassSVM ... done.
- plot coastlines from coverage
- predict species distribution
Area under the ROC curve : 0.993919
time elapsed: 6.31s
# Authors: The scikit-learn developers
# SPDX-License-Identifier: BSD-3-Clause
from time import time
import matplotlib.pyplot as plt
import numpy as np
from sklearn import metrics, svm
from sklearn.datasets import fetch_species_distributions
from sklearn.utils import Bunch
# if basemap is available, we'll use it.
# otherwise, we'll improvise later...
try:
from mpl_toolkits.basemap import Basemap
basemap = True
except ImportError:
basemap = False
def construct_grids(batch):
"""Construct the map grid from the batch object
Parameters
----------
batch : Batch object
The object returned by :func:`fetch_species_distributions`
Returns
-------
(xgrid, ygrid) : 1-D arrays
The grid corresponding to the values in batch.coverages
"""
# x,y coordinates for corner cells
xmin = batch.x_left_lower_corner + batch.grid_size
xmax = xmin + (batch.Nx * batch.grid_size)
ymin = batch.y_left_lower_corner + batch.grid_size
ymax = ymin + (batch.Ny * batch.grid_size)
# x coordinates of the grid cells
xgrid = np.arange(xmin, xmax, batch.grid_size)
# y coordinates of the grid cells
ygrid = np.arange(ymin, ymax, batch.grid_size)
return (xgrid, ygrid)
def create_species_bunch(species_name, train, test, coverages, xgrid, ygrid):
"""Create a bunch with information about a particular organism
This will use the test/train record arrays to extract the
data specific to the given species name.
"""
bunch = Bunch(name=" ".join(species_name.split("_")[:2]))
species_name = species_name.encode("ascii")
points = dict(test=test, train=train)
for label, pts in points.items():
# choose points associated with the desired species
pts = pts[pts["species"] == species_name]
bunch["pts_%s" % label] = pts
# determine coverage values for each of the training & testing points
ix = np.searchsorted(xgrid, pts["dd long"])
iy = np.searchsorted(ygrid, pts["dd lat"])
bunch["cov_%s" % label] = coverages[:, -iy, ix].T
return bunch
def plot_species_distribution(
species=("bradypus_variegatus_0", "microryzomys_minutus_0")
):
"""
Plot the species distribution.
"""
if len(species) > 2:
print(
"Note: when more than two species are provided,"
" only the first two will be used"
)
t0 = time()
# Load the compressed data
data = fetch_species_distributions()
# Set up the data grid
xgrid, ygrid = construct_grids(data)
# The grid in x,y coordinates
X, Y = np.meshgrid(xgrid, ygrid[::-1])
# create a bunch for each species
BV_bunch = create_species_bunch(
species[0], data.train, data.test, data.coverages, xgrid, ygrid
)
MM_bunch = create_species_bunch(
species[1], data.train, data.test, data.coverages, xgrid, ygrid
)
# background points (grid coordinates) for evaluation
np.random.seed(13)
background_points = np.c_[
np.random.randint(low=0, high=data.Ny, size=10000),
np.random.randint(low=0, high=data.Nx, size=10000),
].T
# We'll make use of the fact that coverages[6] has measurements at all
# land points. This will help us decide between land and water.
land_reference = data.coverages[6]
# Fit, predict, and plot for each species.
for i, species in enumerate([BV_bunch, MM_bunch]):
print("_" * 80)
print("Modeling distribution of species '%s'" % species.name)
# Standardize features
mean = species.cov_train.mean(axis=0)
std = species.cov_train.std(axis=0)
train_cover_std = (species.cov_train - mean) / std
# Fit OneClassSVM
print(" - fit OneClassSVM ... ", end="")
clf = svm.OneClassSVM(nu=0.1, kernel="rbf", gamma=0.5)
clf.fit(train_cover_std)
print("done.")
# Plot map of South America
plt.subplot(1, 2, i + 1)
if basemap:
print(" - plot coastlines using basemap")
m = Basemap(
projection="cyl",
llcrnrlat=Y.min(),
urcrnrlat=Y.max(),
llcrnrlon=X.min(),
urcrnrlon=X.max(),
resolution="c",
)
m.drawcoastlines()
m.drawcountries()
else:
print(" - plot coastlines from coverage")
plt.contour(
X, Y, land_reference, levels=[-9998], colors="k", linestyles="solid"
)
plt.xticks([])
plt.yticks([])
print(" - predict species distribution")
# Predict species distribution using the training data
Z = np.ones((data.Ny, data.Nx), dtype=np.float64)
# We'll predict only for the land points.
idx = np.where(land_reference > -9999)
coverages_land = data.coverages[:, idx[0], idx[1]].T
pred = clf.decision_function((coverages_land - mean) / std)
Z *= pred.min()
Z[idx[0], idx[1]] = pred
levels = np.linspace(Z.min(), Z.max(), 25)
Z[land_reference == -9999] = -9999
# plot contours of the prediction
plt.contourf(X, Y, Z, levels=levels, cmap=plt.cm.Reds)
plt.colorbar(format="%.2f")
# scatter training/testing points
plt.scatter(
species.pts_train["dd long"],
species.pts_train["dd lat"],
s=2**2,
c="black",
marker="^",
label="train",
)
plt.scatter(
species.pts_test["dd long"],
species.pts_test["dd lat"],
s=2**2,
c="black",
marker="x",
label="test",
)
plt.legend()
plt.title(species.name)
plt.axis("equal")
# Compute AUC with regards to background points
pred_background = Z[background_points[0], background_points[1]]
pred_test = clf.decision_function((species.cov_test - mean) / std)
scores = np.r_[pred_test, pred_background]
y = np.r_[np.ones(pred_test.shape), np.zeros(pred_background.shape)]
fpr, tpr, thresholds = metrics.roc_curve(y, scores)
roc_auc = metrics.auc(fpr, tpr)
plt.text(-35, -70, "AUC: %.3f" % roc_auc, ha="right")
print("\n Area under the ROC curve : %f" % roc_auc)
print("\ntime elapsed: %.2fs" % (time() - t0))
plot_species_distribution()
plt.show()
腳本的總執行時間: (0 分鐘 6.454 秒)
相關範例